Among these, 19 species are widely distributed in the Oriental Region (Guizhou, Sichuan, Hunan, Hubei, Guangxi, Guangdong, Fujian, and Zhejiang), and only A. 
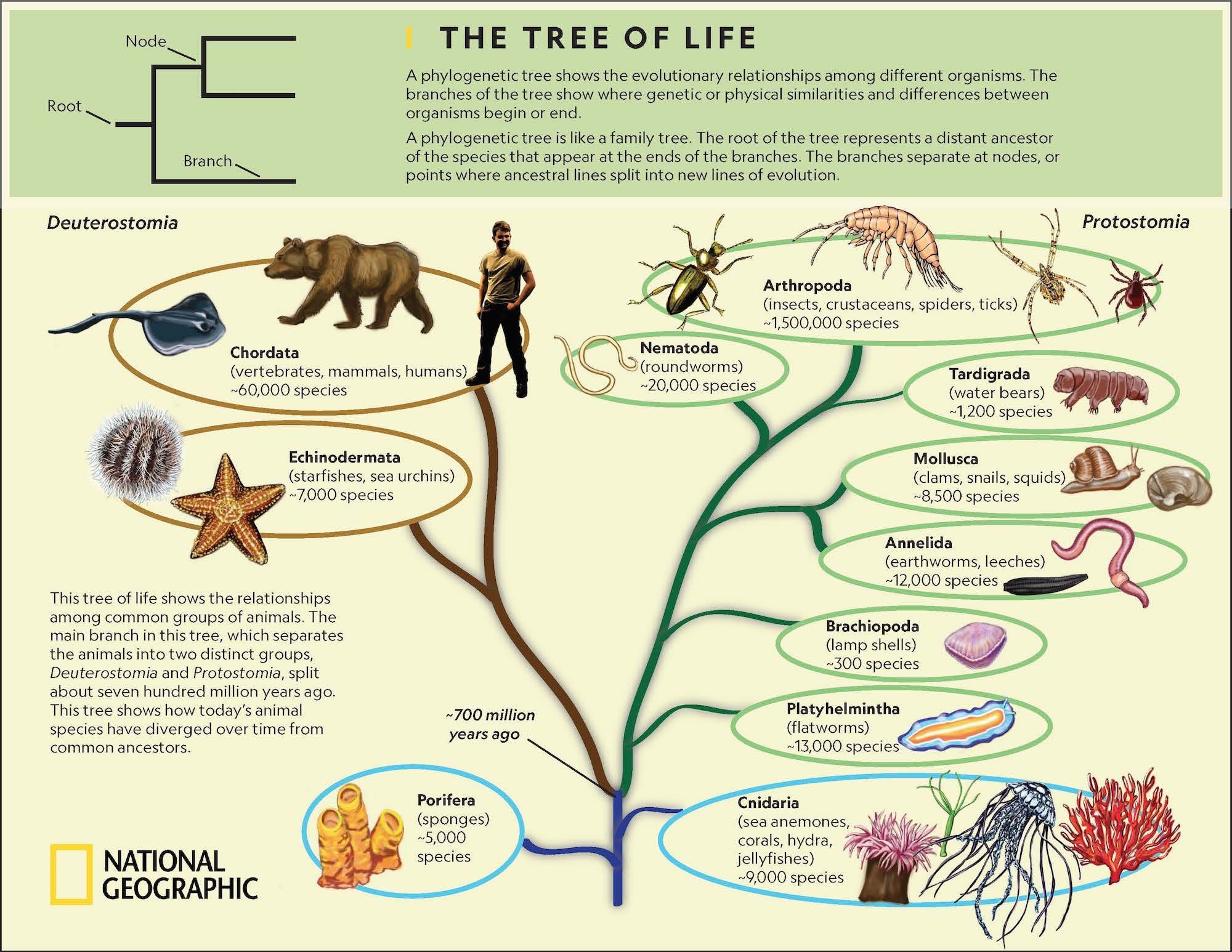
So far, Abrus is only restricted to China with 19 valid described species, which are quite similar in body coloration and difficult to distinguish based on external morphology, but the male genitalia with a unique structure of aedeagus are considerably differences among these species. , Yang & Chen, and Xing & Li further added 13 new species from the Oriental and Palaearctic parts of China. After that, Li & Wang, Dai & Zhang, Li et al. hengshanensis Dai & Zhang, 2002) from China. The leafhopper genus Abrus belonging to the tribe Athysanini of subfamily Deltocephalinae, was originally described by Dai & Zhang with six new species (type species: A. Due to their high genome copy numbers, multiple genome-level characteristics, relatively high evolutionary rate, and greater phylogenetic informativeness than single mitochondrial genes, mitogenome sequences have been widely used in various phylogenomic studies. The representative mitochondrial (mt) genomes of almost all insect orders from higher-level to lower taxonomic ranks have extensively been utilized for studying phylogeny, population genetics, comparative and evolutionary genomics, molecular evolution, and identification at various taxonomic levels. Insect mitochondrial genome is a small double-stranded circular molecule with remarkable conservation in size ranging from 14–20 kb that encodes 37 genes: 13 protein-coding genes, two ribosomal RNAs, 22 transfer RNAs genes, and a non-coding A + T-rich region (or control region). Phylogenetic trees reconstructed herein based on the BI and ML analyses consistently recovered monophylitic Athysanini, except Watanabella graminea (Athysanini) in Opsiini with high support values. ConclusionĪt present, Abrus belongs to the tribe Athysanini based on both morphological and molecular datasets, which is strongly supported in present phylogenetic analyses in both BI and ML methods using the six concatenated datasets: amino acid sequences and nucleotides from different combinations of protein-coding genes (PCGs), ribosomal RNA (rRNAs), and transfer RNA (tRNAs). Phylogenetic analyses based on 13 PCGs, 2 rRNA genes, and 22 tRNA genes consistently recovered the monophyletic Opsiini, Penthimiini, Selenocephalini, Scaphoideini, and Athysanini (except Watanabella graminea, previously sequenced species as Chlorotettix nigromaculatus) based on limited available mitogenome sequence data of 37 species. All 22 tRNA genes had typical cloverleaf secondary structures, except for trnS1 (AGN) which lacks the dihydrouridine arm, and distinctively trnG in the mitogenome of A. The nucleotide composition of these genomes is highly biased toward A + T nucleotides (76.2%, 76.3%, and 74.7%), AT-skews (0.091 to 0.095, and 0.095), negative GC-skews (− 0.138, − 0.161, and − 0.138), and codon usage. These Abrus species shared highly conserved mitogenomes with similar gene order to that of the putative ancestral insect with 37 typical genes and a non-coding A + T-rich region. yunshanensis Chen, Yang & Li, 2012 (15,768bp) (Athysanini), and compared with published mitogenome sequence of A. In this study, we sequenced and analyzed the complete mitochondrial genomes (mitogenomes) of Abrus daozhenensis Chen, Yang & Li, 2012 (16,391bp) and A. Although the taxonomy of Abrus are well updated, the references on comparative mitogenomic analyses of Abrus species are only known for a single species. The bamboo-feeding leafhopper genus Abrus belong to the tribe Athysanini of subfamily Deltocephalinae, which currently comprises 19 valid described species, and are limited to the Oriental and Palaearctic regions in China. The phylogenetic position and classification of Athysanini are poorly defined, as it includes a large group of polyphyletic genera that have historically been assigned to it mainly because they still exhibit the most typical deltocephaline genitalic and external body characters but lack the distinctive characteristics that other tribes possess.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
